 Loading... Please wait...
Loading... Please wait...Categories
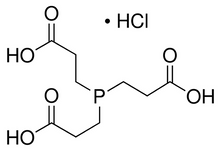
TCEP HCl
(TCEP; TCEP HCl; Tris (2-Carboxyethyl) phosphine Hydrochloride)
Purity > 98%
| CAS Number: |
51805-45-9
|
| Chemical Formula: |
C9H16O6PCl
|
| Molecular Weight: |
286.6 g/mol
|
TCEP HCl is a reducing agent suitable for protein applications. TCEP is often used for the reduction of disulfide bonds within proteins. This versatile compound is very soluble in aqueous solutions (310 grams/L in water). TCEP is stable in both acidic and basic solutions (pH 1.5 to 8.5).
TCEP has many other advantages such as:
- TCEP HCl is stable in air.
- TCEP HCl is odorless eliminating the need to preform reactions within a fume hood.
- TCEP HCl is efficient and can be used to reduce disulfide bonds at low concentrations (5 mM).
- TCEP HCl may be used as a substitute for DTT (dithiothreitol) at a final concentration of 50 mM.
Store: Below 4 °C
Free Shipping within the Continental USA
TCEP Certification of Analysis
Manufacture and Expiration Data available upon request.
| Test | Speficiations | Results |
| Appearance | Off-white crystalline powder | Corresponds |
| Solution (1% in Water) | Colorless, clear | Colorless, clear |
| Loss on Drying | Maximum 1.0 % | Maximum 0.3 % |
| Melting Point | 181 °C ± 2 °C | 179 °C |
| Assay (IC) | Min. 98.0% | 99.6 % |
| FTIR | Reference spectrum | Corresponds |
Preparation of 0.5 M TCEP Stock Solution
(Note: TCEP solutions are commonly neutralized due to having a pH of ~2.5 in water.)
- Add 5.733 g of TCEP to a 50 ml conical vial, making sure to get as much of the TCEP as possible into the vial.
- Add 35 ml of cold molecular biology grade water to the vial, and dissolve the TCEP. This resulting solution is acidic, with an approximate pH of 2.5.
- Bring the solution to pH 7.0 with 10N NaOH or 10N KOH.
- Bring the resulting solution to 40 ml with molecular biology grade water.
- Aliquot into 1.6 ml Microcentrifuge tubes and store at -20 °C.
View this method on protocols.io and use it directly at your bench
Additional Benefits of TCEP HCl
- TCEP is soluble in a many aqueous buffers and minimally in organic solvents such as methanol or ethanol.
- TCEP is stable over a wide range of pH levels
- TCEP can be prepared in 10X stock solutions
- While TCEP can be used in Phosphate buffers, it has the tendency to be unstable around neutral pH levels. If the TCEP is needed in the buffer, prepare it immediately before use.
- A 50mM TCEP solution may be substituted for DTT or β-mercaptoethanol (2-ME) in sample loading buffer for SDS-PAGE
- TCEP should not be used for isoelectric focusing, due to its charge once in solution.
Trypsin Digestion Protocol for a 300 μL Protein Mixture (e.g. cell lysates)
- Add 108 mg of 6 M urea to 300 μL of 50 mM ammonium bicarbonate containing protein.
- Add 20 μL of 1.5 M Tris pH 8.8.
- Add 7.5 μL of 200 mM TCEP and incubate for 1 hr at 37 °C.
- Add 60 μL of 200 mM iodoacetamide.
- Vortex and let sit for 1 hr at room temperature in a dark area.
- Add 60 μL of 200 mM DTT (dithiothreitol)
- Vortex and let sit for 1 hr at room temperature.
- Aliquot 150 μL into each of the 31.5 mL eppendorf tubes.
- Add 800 μL of 25 mM NH4HCO3 to each tube to dilute the urea.
- Add 200 μL of MeOH to each tube.
- Add Trypsin to protein at a ratio of 1:50.
- Incubate overnight at room temperature.
- Speedvac to dryness.
- Add 200 μL of molecular biology grade water to tubes three times and speedvac to reduce NHH2CO3.
- Store at -80 °C.
Adapted from the Goodlett Lab of the University of Washington
Comparable Items:
| Acros Organics | 36383-0010 |
| Alfa Aesar | 40587-04 |
| Alfa Aesar | 40587-02 |
| Alfa Aesar | 40587-09 |
| Alfa Aesar | H51864-AC |
| Alfa Aesar | H51864-AA |
| Alfa Aesar | H51864-AE |
| Amresco | K831-2G |
| Amresco | K831-10G |
| Anaspec Inc | 20490 |
| Anaspec Inc | 20290 |
| Anaspec Inc | 22582 |
| Bel-Art | 85178 |
| Boekel | 29700 |
| EMD Millipore | 4.85079.0002 |
| EMD Millipore | 580560-1GM |
| Electron Microscopy Science | 77720 |
| Keystone | TS-48927 |
| Keystone | TS-48920 |
| Lagasse | 35602 |
| Quanta BioDesign | 10014-1000MG |
| Quanta BioDesign | 10014-5G |
| Quanta BioDesign | 10014-25G |
| Restek | 23011 |
| Restek | 20504 |
| Safety & Industrial Supplies | 23031 |
| Strem | 15-7400.5 |
| Strem | 15-7400.1 |
| TCI America | T1656-25G |
| TCI America | T1656-5G |
| TCI America | T1656-1G |
| Fisher Scientific | 24500 |
| Fisher Scientific | 20907 |
| Fisher Scientific | 35603 |
| Fisher Scientific | 24110 |
| Fisher Scientific | 24115 |
| Fisher Scientific | 24510 |
| Fisher Scientific | 26101 |
| Fisher Scientific | 26103 |
| Fisher Scientific | 44889 |
| Fisher Scientific | 77712 |
| Fisher Scientific | 24308 |
| Fisher Scientific | 28905 |
| Fisher Scientific | 20291 |
| Fisher Scientific | 23030 |
| Fisher Scientific | 26777 |
| Fisher Scientific | 20408 |
| Fisher Scientific | 20491 |
Celebrity Endorsements
Burns, et al. (1991) Selective reduction of disulfides by tris(2-carboxyethyl)phosphine J. Org. Chem., 56 (8), 2648–2650.
Description: Tris(2-carboxyethy1)phosphine (TCEP) reduces disulfides rapidly and completely in water at pH 4.5. TCEP displays a preference for more strained disulfide bonds and follows the usual mechanism for the reduction of disfulfides by phosphines in water. A method of large scale production of TCEP is also described via acidic hydrolysis.
Getz, et al. (1999) A Comparison between the Sulfhydryl Reductants Tris(2-carboxyethyl)phosphine and Dithiothreitol for Use in Protein Biochemistry Analytical Biochemistry 273, 73–80.
Decription: Comparing and contrasting of TCEP with DTT for the use in protein biochemistry. TCEP has been shown to be significantly more stable than DTT at pH values above 7.5, and as a faster and stronger reductant than DTT at pH values below 8.0. Thus TCEP is a useful reductant over a much wider pH range (1.5–8.5) than DTT, although the buffer composition, including the presence of phosphates, can adversly affect TCEP stability. In addition, TCEP has been advertised as being unreactive with thiol-reactive compounds, thereby eliminating the need to remove it before certain labeling applications.
Results were shown that during protein purification, Ni concentrations contaminating proteins eluted from Nickel affinity columns cause rapid oxidation of DTT without affecting TCEP. For long-term storage of proteins, TCEP is significantly more stable than DTT without metal chelates such as EGTA in the buffer, whereas DTT is more stable if metal chelates are present. In electron paramagnetic resonance (EPR) spectroscopy utilizing nitroxide spin labels, TCEP is highly advantageous: spin labels are two to four times more stable in TCEP than DTT. Neither reducing agent had a significant effect at 0.1 mM concentrations iodoacetamide attachment to the protein of interest (myosin). The reductants equally preserve myosin’s enzymatic activity, which is sensitive to sulfhydryl oxidation. Thus TCEP has advantages over DTT, although the choice of reductant is application specific. The paper also includes a look at long term stability varying temperature as well as ions (Fe, Mg, EGTA) present.
Han, J.C. and Han, G. Y. (1994) A Procedure for Quantitative Determination of Tris(2-Carboxyethyl)phosphine, an Odorless Reducing Agent More Stable and Effective Than Dithiothreitol Analytical Biochemistry, 220 (1), 5-10.
Krezel, et al. (2003) Coordination properties of tris (2-carboxyethyl) phosphine, a newly introduced thiol reductant, and its oxide. Inorg. Chem., 42 (6), 1994–2003.
Cline, et al. (2004) New water-soluble phosphines as reductants of peptide and protein disulfide bonds: reactivity and membrane permeability. Biochemistry, 43 (48), 15195–15203.
Applications of TCEP in Protein Crystallization:
Jancarik, J., et. al. (2004) Optimum solubility (OS) screening: an efficient method to optimize buffer conditions for homogeneity and crystallization of proteins. Acta Cryst. D. 60. 1670-1673.
Page, R., et. al. (2004) Crystallization data mining in structural genomics: using positive and negative results to optimize protein crystallization screens. Methods in Macromolecular Crystallization . 34. 373-389.
Description: Tris(2-carboxyethy1)phosphine (TCEP) reduces disulfides rapidly and completely in water at pH 4.5. TCEP displays a preference for more strained disulfide bonds and follows the usual mechanism for the reduction of disfulfides by phosphines in water. A method of large scale production of TCEP is also described via acidic hydrolysis.
Getz, et al. (1999) A Comparison between the Sulfhydryl Reductants Tris(2-carboxyethyl)phosphine and Dithiothreitol for Use in Protein Biochemistry Analytical Biochemistry 273, 73–80.
Decription: Comparing and contrasting of TCEP with DTT for the use in protein biochemistry. TCEP has been shown to be significantly more stable than DTT at pH values above 7.5, and as a faster and stronger reductant than DTT at pH values below 8.0. Thus TCEP is a useful reductant over a much wider pH range (1.5–8.5) than DTT, although the buffer composition, including the presence of phosphates, can adversly affect TCEP stability. In addition, TCEP has been advertised as being unreactive with thiol-reactive compounds, thereby eliminating the need to remove it before certain labeling applications.
Results were shown that during protein purification, Ni concentrations contaminating proteins eluted from Nickel affinity columns cause rapid oxidation of DTT without affecting TCEP. For long-term storage of proteins, TCEP is significantly more stable than DTT without metal chelates such as EGTA in the buffer, whereas DTT is more stable if metal chelates are present. In electron paramagnetic resonance (EPR) spectroscopy utilizing nitroxide spin labels, TCEP is highly advantageous: spin labels are two to four times more stable in TCEP than DTT. Neither reducing agent had a significant effect at 0.1 mM concentrations iodoacetamide attachment to the protein of interest (myosin). The reductants equally preserve myosin’s enzymatic activity, which is sensitive to sulfhydryl oxidation. Thus TCEP has advantages over DTT, although the choice of reductant is application specific. The paper also includes a look at long term stability varying temperature as well as ions (Fe, Mg, EGTA) present.
Han, J.C. and Han, G. Y. (1994) A Procedure for Quantitative Determination of Tris(2-Carboxyethyl)phosphine, an Odorless Reducing Agent More Stable and Effective Than Dithiothreitol Analytical Biochemistry, 220 (1), 5-10.
Krezel, et al. (2003) Coordination properties of tris (2-carboxyethyl) phosphine, a newly introduced thiol reductant, and its oxide. Inorg. Chem., 42 (6), 1994–2003.
Cline, et al. (2004) New water-soluble phosphines as reductants of peptide and protein disulfide bonds: reactivity and membrane permeability. Biochemistry, 43 (48), 15195–15203.
Applications of TCEP in Protein Crystallization:
Jancarik, J., et. al. (2004) Optimum solubility (OS) screening: an efficient method to optimize buffer conditions for homogeneity and crystallization of proteins. Acta Cryst. D. 60. 1670-1673.
Page, R., et. al. (2004) Crystallization data mining in structural genomics: using positive and negative results to optimize protein crystallization screens. Methods in Macromolecular Crystallization . 34. 373-389.